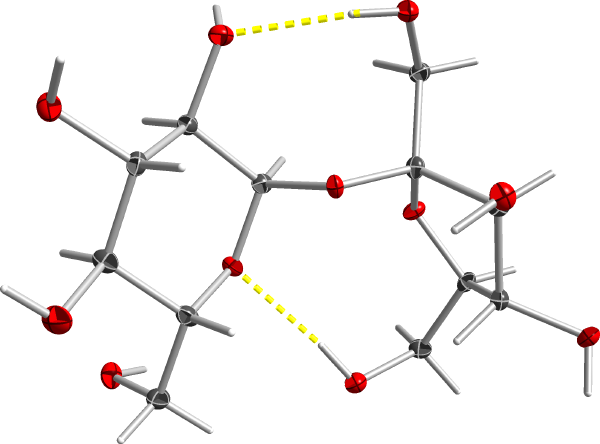
Routine structure determination with SHELX: sucrose
Note: This tutorial uses older programs: shelxs, shelxl97, and shelxtl XP.This tutorial is intended to guide you step-by-step through a routine structure determination. It is intentionally very simplistic. In spite of what you may have been told, many structures are quite straightforward. You simply have to know what to do, which buttons to click, what to type, as well as when and how to make decisions. The structure in question is sucrose.