4/6) KappaCCD: Scaling with SADABS
We're now ready to run separate SADABS jobs for the 'fast' and 'slow' scans.
Here, we'll process the 'fast' scan first, but the order doesn't matter. In a terminal
window, start SADABS like this:
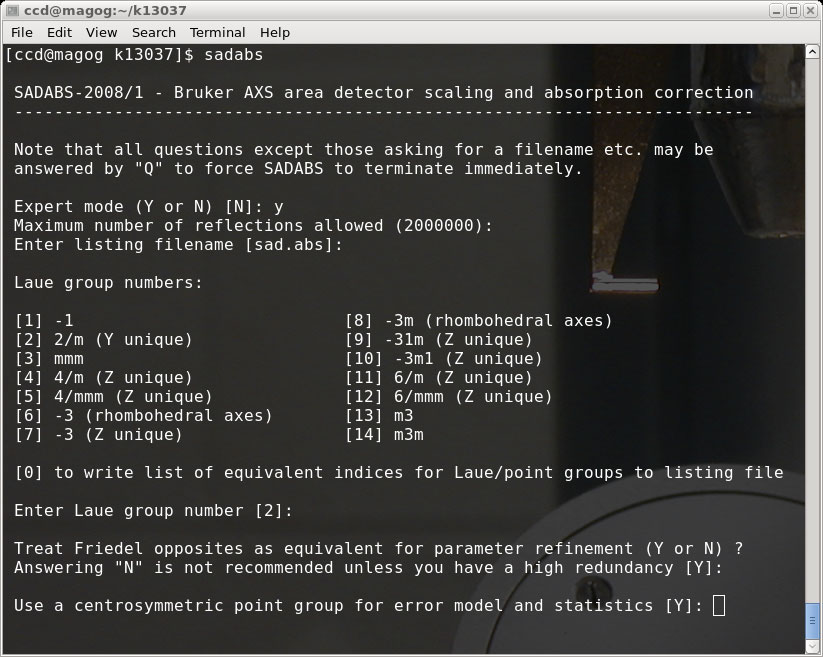
This is not intended to be a tutorial on how to use SADABS, but some
information about the runs for this particular sample is given in the pictures.
Different crystals need different treatment and different parameters. It is, of
course, important to get the Laue group right (or at the very least to not use a Laue
group of too high symmetry!).
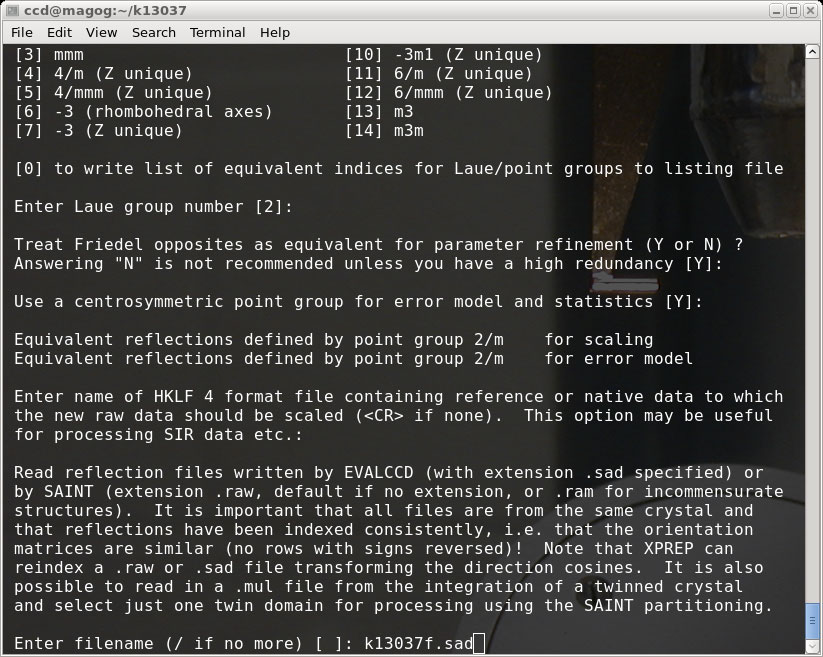
When prompted for the filename, enter the first (here it's the 'fast') .sad file:
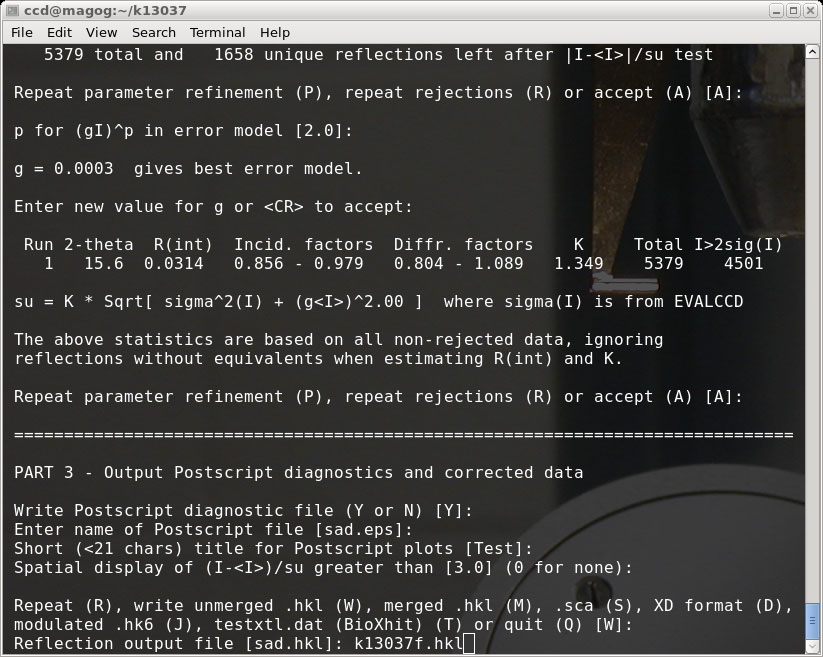
Some steps in the SADABS process have been omitted in the images - just be sure
to use sensible choices for your crystal! It makes sense to use the same 'fast'
(f) and 'slow' (s) designators for the output '.hkl' files (as in the above picture).
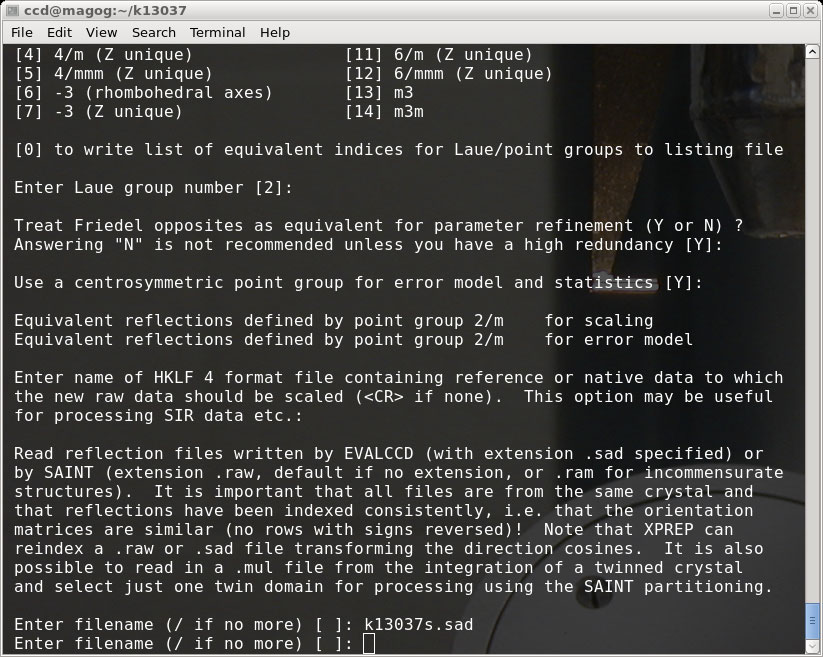
Repeat the process for the other scan. Since it is the same crystal you should use the
same input parameters (other than file name!).
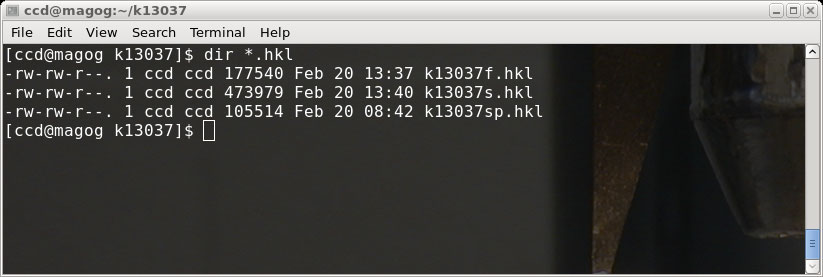
The end result of these separate SADABS runs is a pair of '.hkl' files for the
'fast' and 'slow' scans. In the above picture you can also see a file 'k13037sp.hkl',
which is the file from the Scalepack run (made for comparison purposes).
Notice that its file size is smaller than either of the SADABS generated
files. The reason is that Scalepack merged equivalent reflections. We will
not use this file here, but it was comparison of different data treatment methods
that led to the procedure described here. The next step is to find and eliminate
overloads from the 'slow' dataset (part 5).
1) Extract the intensity data from the diffraction frames.
2) Run a first pass through with Scalepack.
3) Create '.sad' input files for SADABS.
4) Run SADABS separately for fast and slow scans.
5) Identify overload intensities and edit slow .hkl file.
6) Merge fast and slow scans in XPREP.
2) Run a first pass through with Scalepack.
3) Create '.sad' input files for SADABS.
4) Run SADABS separately for fast and slow scans.
5) Identify overload intensities and edit slow .hkl file.
6) Merge fast and slow scans in XPREP.