3) Solve the structure
You are now ready to run one of the structure solving routines that are present in the maXus package. From the maXus menu:
You should click the "Solve" menu item. A sub-menu will appear with a few choices. Generally the fastest one of these structure solution algorithms, and the one that gives perhaps the"cleanest" solution (provided you gave it a formula that was close to correct) is the first default of SIR92. The other SIR options are slower and not always any better. SHELXS is also available - if SIR92 fails you might try SHELXS next).
Anyway, assuming you choose SIR92, you will get a window like this to show that everything is going according to plan:
After the phasing and Fourier re-building processes are complete, you will get a graphical rendition of your structure.
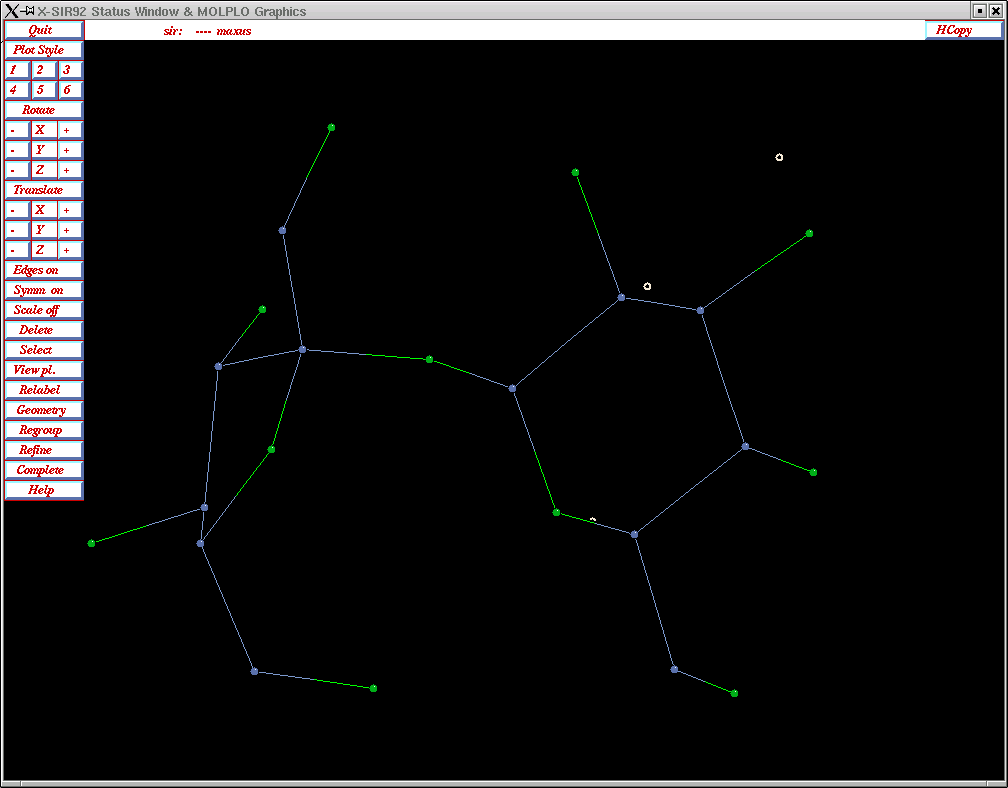
SIR92 is pretty good at assigning atom types provided it has a formula that is approximately correct. In the example here it is pretty easy to pick out the sucrose molecule. You can spin it around to inspect it here, but there are better molecular editing features available elsewhere on the maXus menu. In the top left corner, click the "Quit" button.
This will give you a summary of the structure solution run, like this:
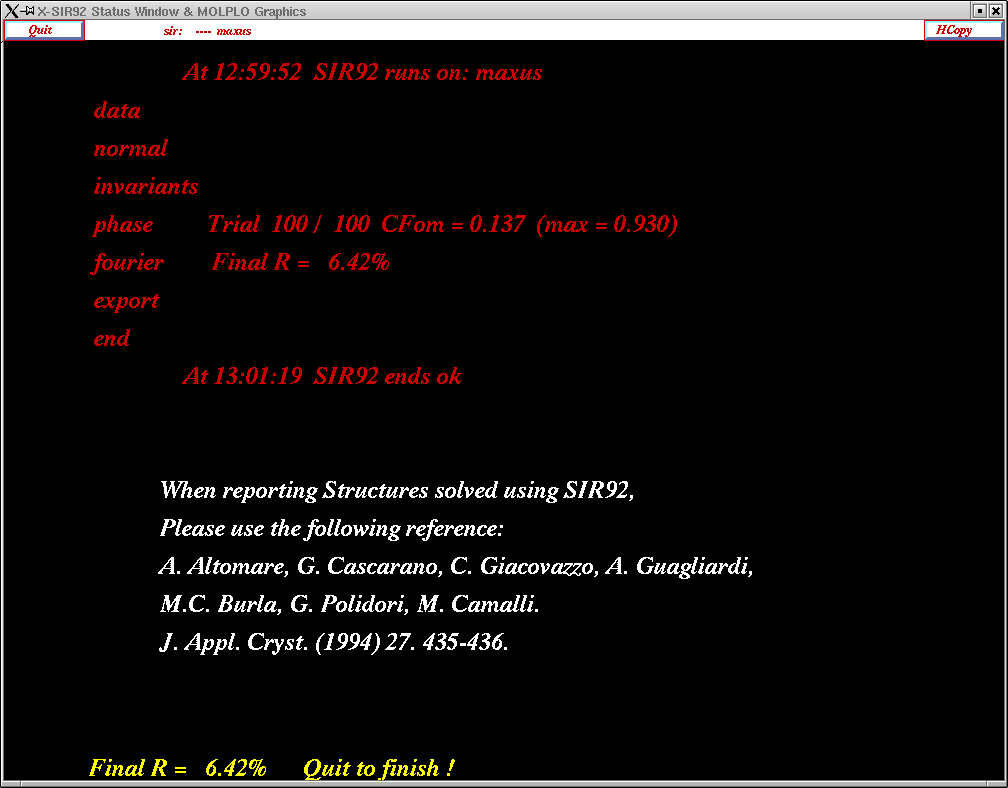
You need to click "Quit" again ...
... and click "OK" on this window to get back to the main maXus window:

Provided that everything went well, your structure is now "solved". It is not yet complete, however. In fact, it may be nowhere near complete. It is very likely that many of the atom types are specified incorrectly. There will likely be some atoms missing and there will probably be some spurious "atoms". The model needs to be edited before beginning refinement. This can be done from within the "Model" sub-program. This is covered briefly in the next installement of this tutorial.
Go on to maXus structure solution guide section 4
Go back to maXus structure solution guide section 2
Go back to maXus structure solution guide main menu
Return to the main Tutorials page or to the main X-Ray Lab page