Introduction
PrimeXP [python retro interactive model editor (inspired by) XP ] is an interactive crystal structure editor with some intentional resemblance to the program that inspired it, George Sheldrick's XP. It includes many of the same four-letter commands as XP, though some have slightly altered and/or expanded functionality. There are also a few new commands that serve as enhancements or extensions. In this capacity, PrimeXP bridges the gap between the crystallographic refinement engine and publication-quality diagrams. It reads and writes SHELXL *.res and *.ins files, plus a few other useful formats. It is not intended as a competitor to programs such as ShelXle or Olex2, but there is some overlap. It is much more closely related to XP. For example, similar to XP, PrimeXP does not spawn the SHELXL refinement program, and probably never will. It does, however, provide a wide array of structure drawing and analysis tools that seek to match or surpass other small-molecule molecular graphics programs.
PrimeXP is built using Python, PyQt6, and PyOpenGL, using NumPy, scikit-image, and SciPy for geometric and topological mathematics, graphics, etc., and is highly customisable. Visualisation is handled by OpenGL 4.1, enabling real-time rendering of features such as atomic displacement ellipsoids, a variety of atom and bond styles, electron density in 2D and 3D, atom label collision avoidance (a never-ending work in progress - a true can-of-worms), etc.
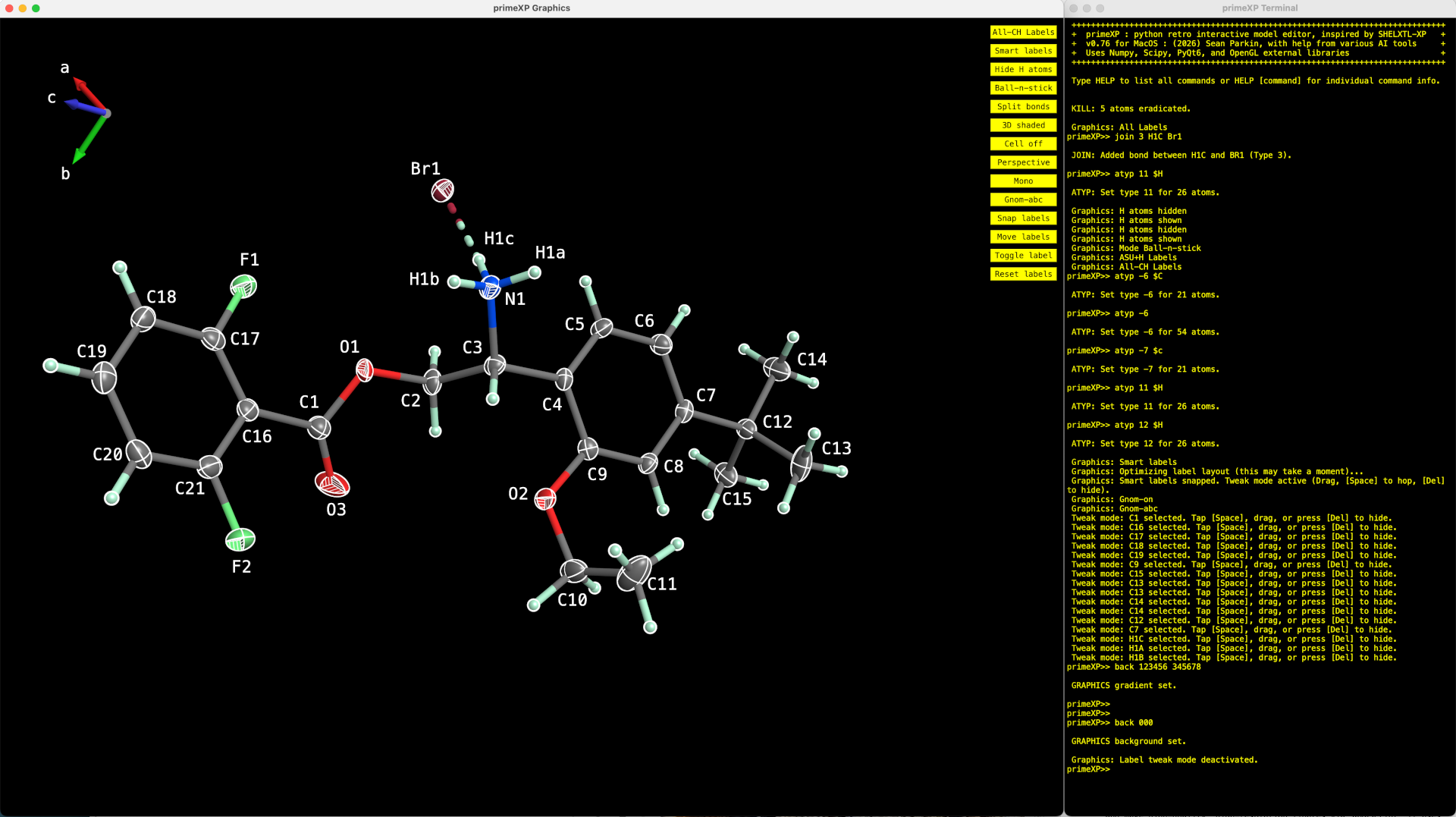
The PrimeXP graphical interface (left) showing a structure with 50% displacement ellipsoids for non-H atoms and the terminal emulator (right). The axis direction gnomon in the upper left has a 3-state toggle (off/on/on+labels). The yellow column of buttons on the right side of the graphics window are an homage to George's XP program. The code is modular, so would be straightforward to re-tool these as rows of drop-down menus, as per more modern GUIs.
PrimeXP Features
The program uses a hybrid interface, combining interactive 3D graphics with a bash-like terminal emulator, which should be familiar to experienced crystallographers. Key features include:
- Advanced rendering: Customisable representations including wireframe, stick, ball-and-stick, anisotropic displacement ellipsoids, with or without major ellipses and/or cutout octants, and Bürgi-style 'peanut' plots.
- Symmetry generation: Real-time application of space-group operators and cell translations to expand the asymmetric unit and examine packing motifs.
- Rigid-body analysis: Translation-Libration-Screw (TLS) dashboard for analyzing rigid molecular fragments.
- Geometric analysis: Tools for calculating mean planes, torsion angles, interactive distance/angle measurements, and Cremer-Pople ring puckering parameters.
- Hydrogen modelling: Automated hydrogen atom addition for various hybridization states (e.g., sp2, sp3) using standard riding model geometries. This is more sophisticated than HADD in XP, but much less so than the HFIX facilities in SHELXL.
- Export: Generation of high-resolution diagrams, orthogonal coordinate files, newly edited *.ins file etc. suitable for publication or further analysis.
A few PrimeXP pictures
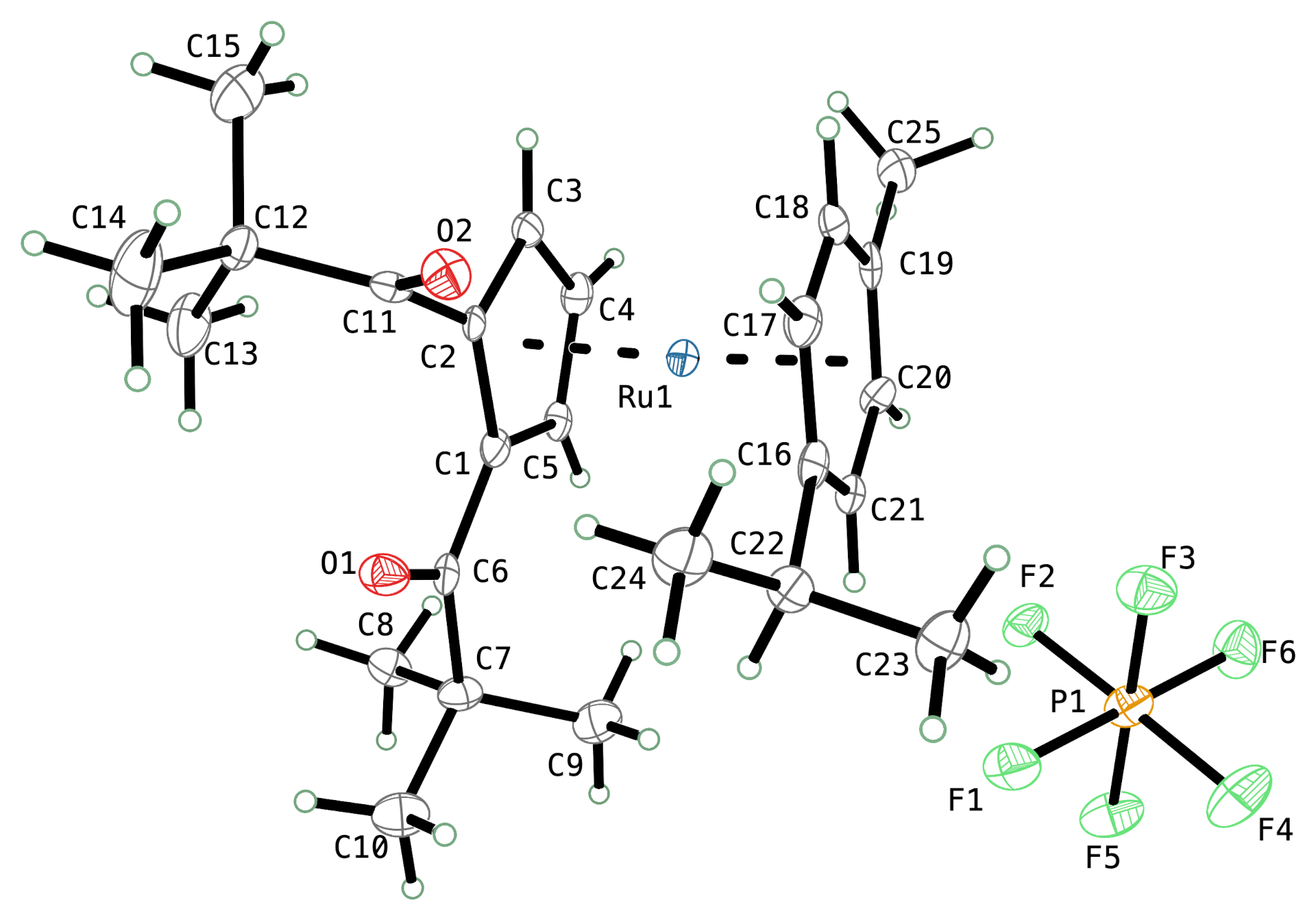
A typical ellipsoid plot in 'TELP' mode from PrimeXP. The program currently has 28 different 'ATYP' atom types for ellipsoids and spheres, with or without various kinds of shading/octant views, and it now includes anisotropic displacement parameters in the style of Hans-Beat Bürgi's PEANUT plots. Unlike XP, TELP in PrimeXP is fully 3D-interactive, as are the 'peanuts' (vide infra ).
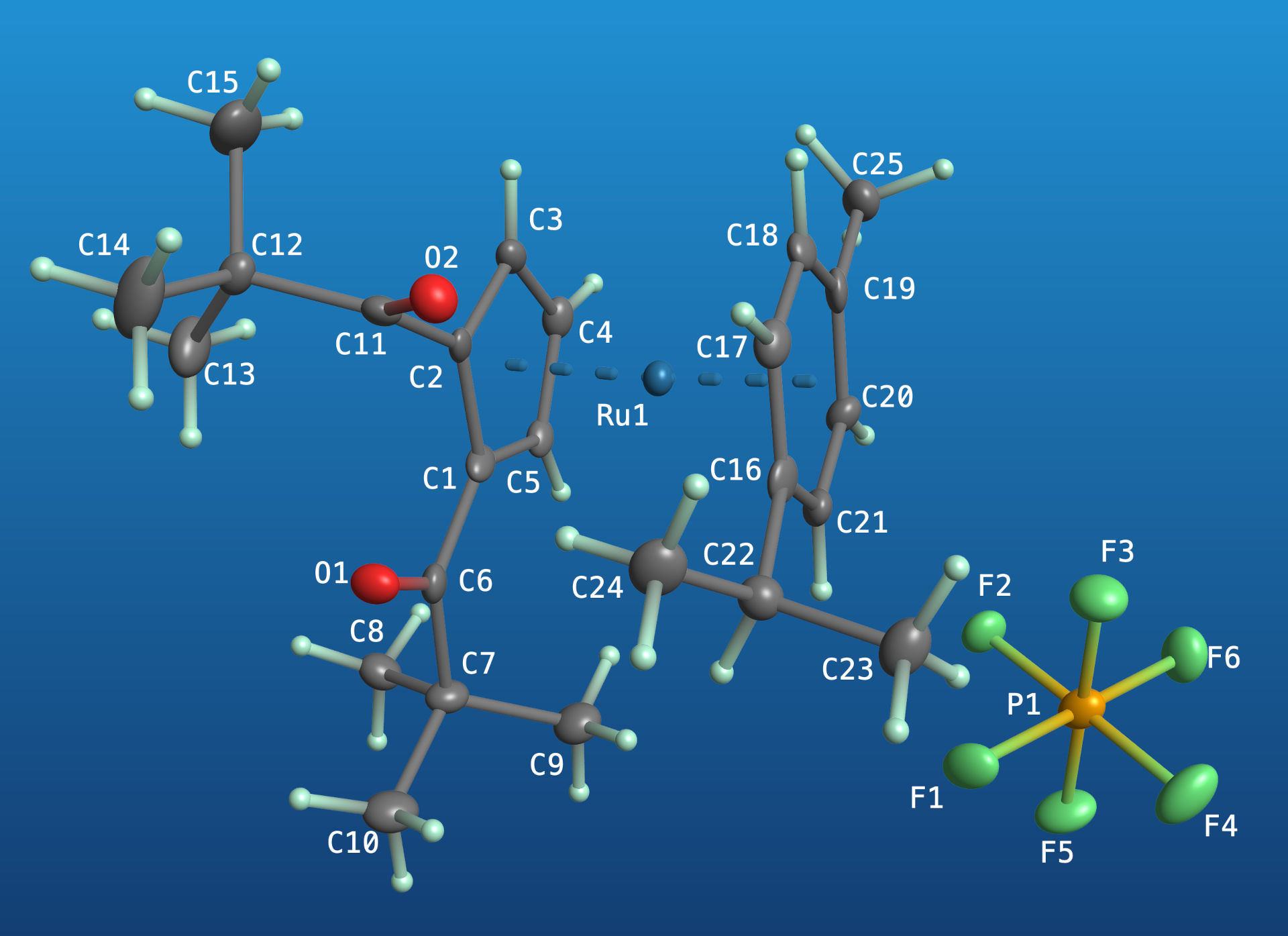
The same molecular orientation, but with 3D-shaded ellipsoids and spheres on a blue gradient background. Colours may be specified using 6-digit hexcodes (~16 million colours), 3-digit hexcodes (~4 thousand colours), or 148 named 'web' colours. The label colours may be set individually (via colour name or hexcode), but default to black or white, automatically switching based on the overall luminosity of the scene.
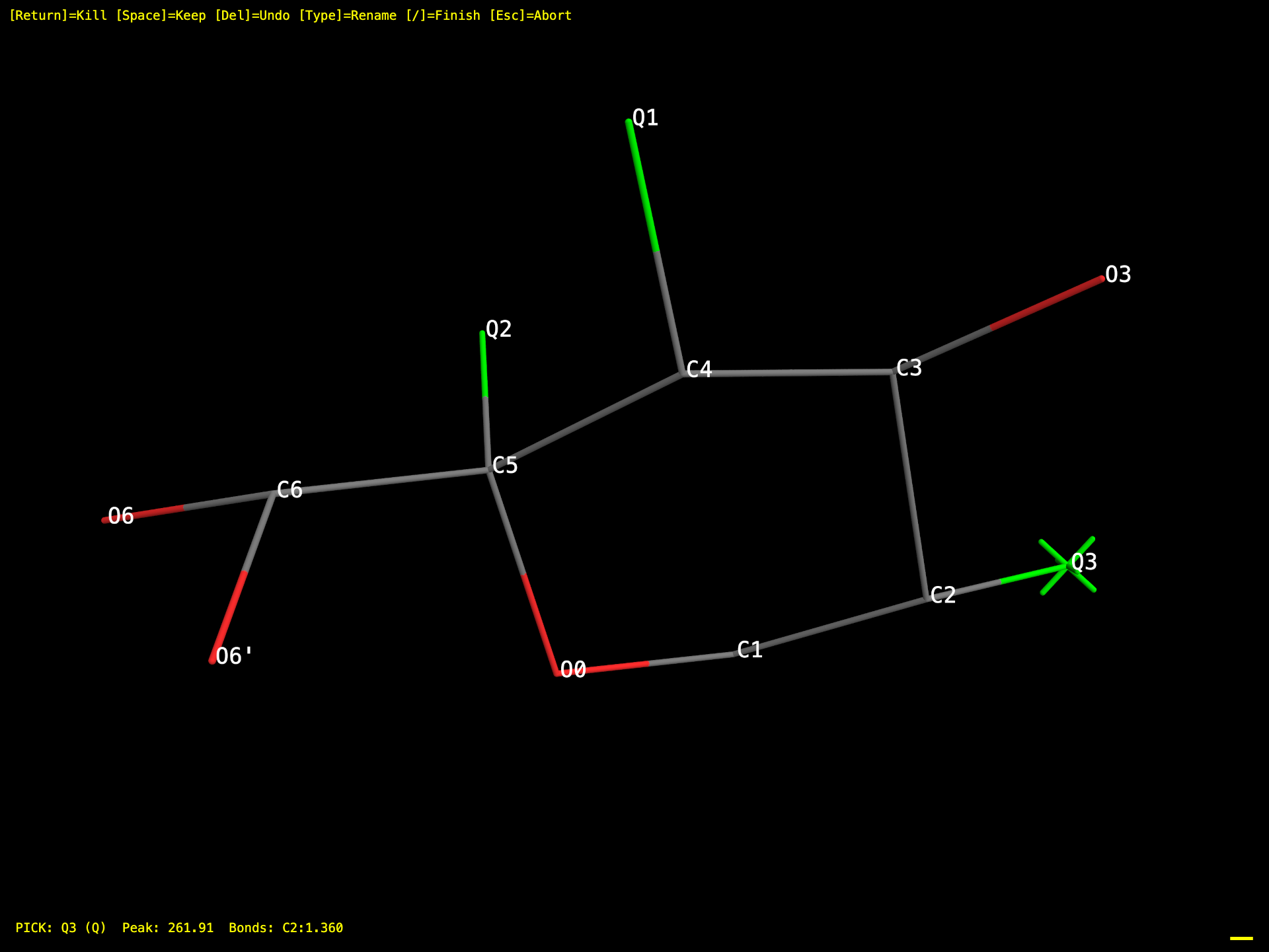
Model editing using PICK is similar to its counterpart in the original XP. In PrimeXP, however, PICK is fully 3D interactive, and the PICK sequence can be interrupted by mouse clicks on any visible Q-peak or atom.
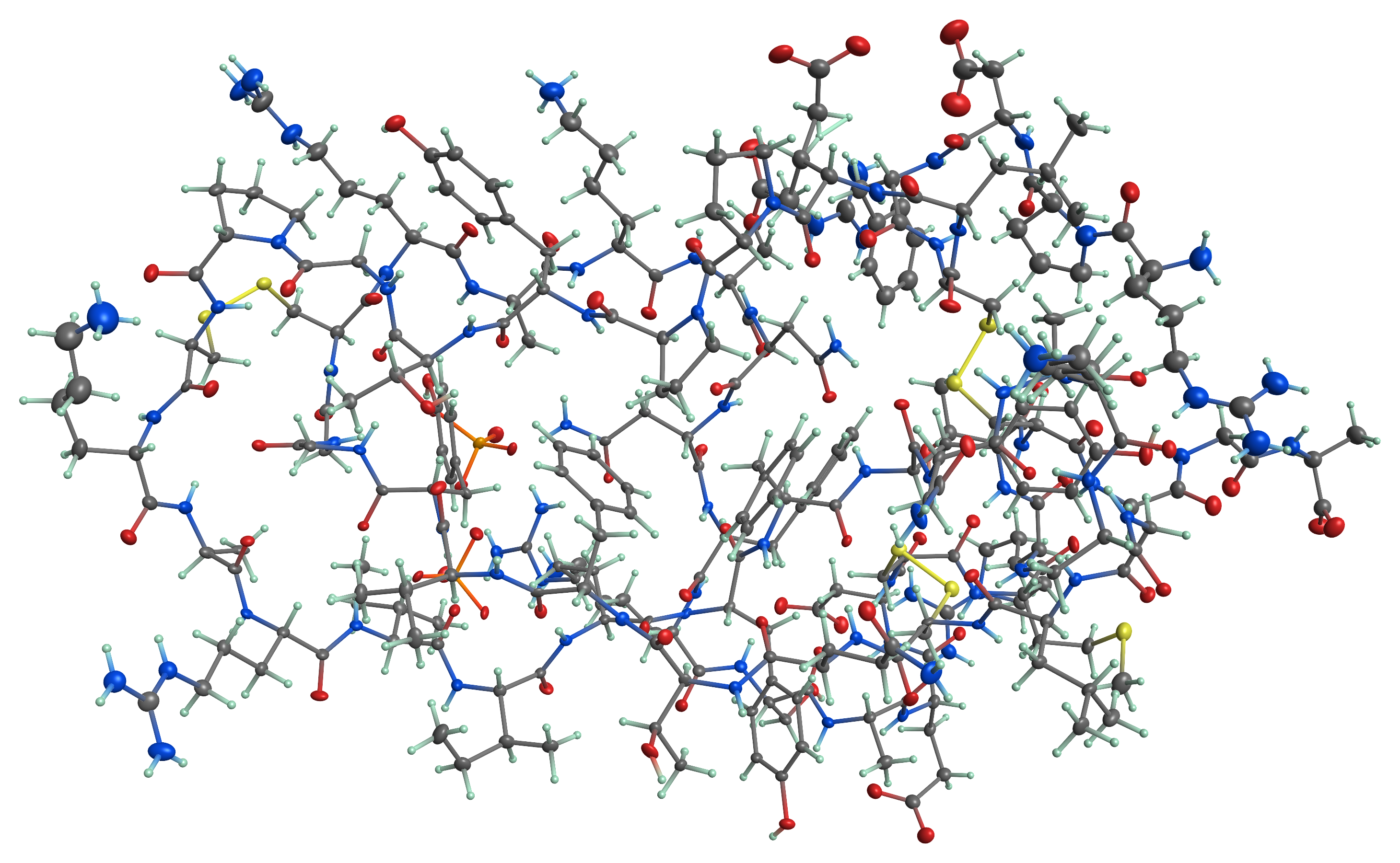
Although it was not written with proteins in mind, PrimeXP is capable of rendering protein structures, such as the above 1.1 Å resolution structure of BPTI.
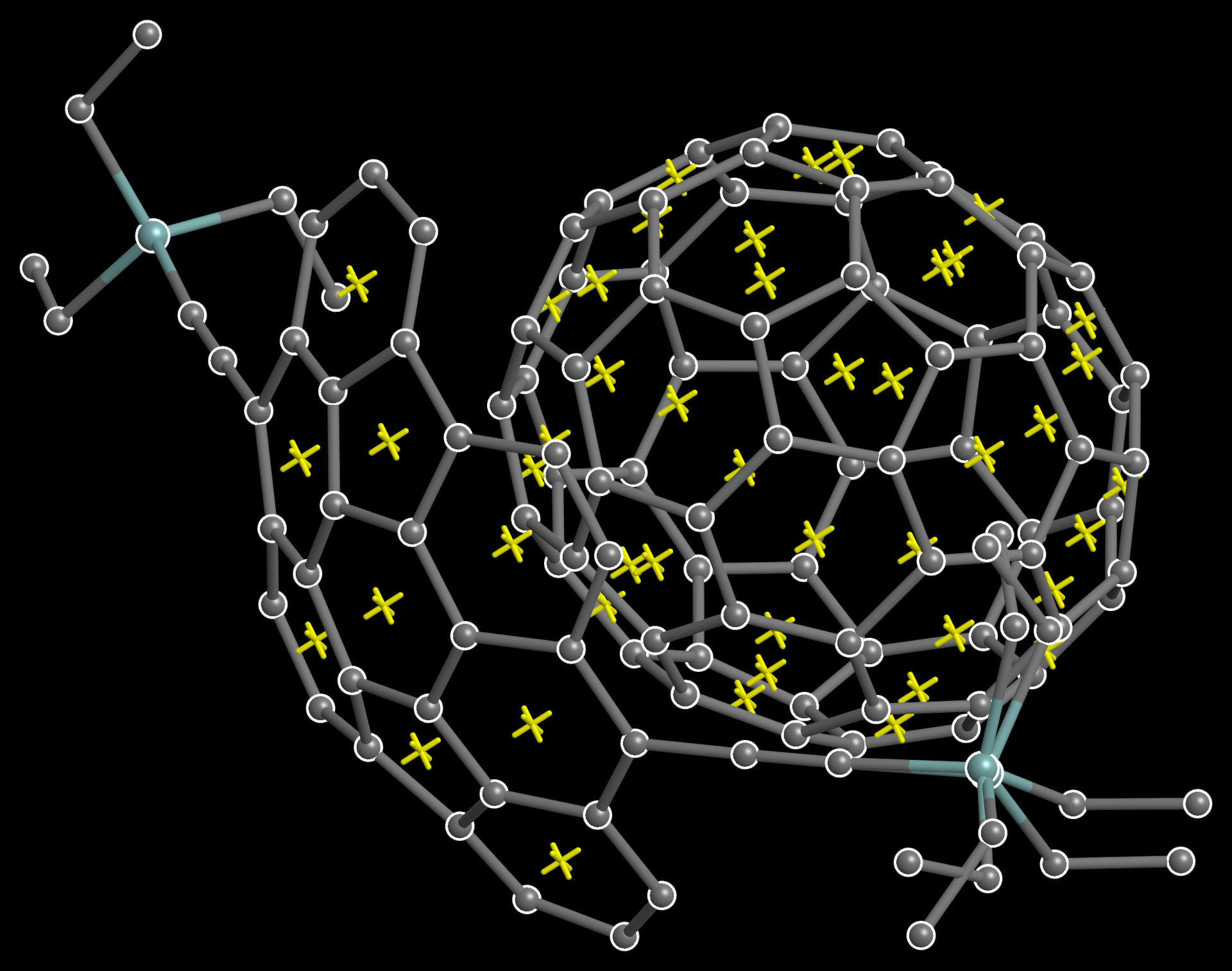
The RING/X command automatically places a dummy 'X' atom at the centroid of each ring (default 3-8 membered rings), as in the above C70 buckyball/buckybowl complex. In contrast to CENT/X in XP, that's 46 dummy atoms placed by a single 'RING/X' command, without having to explicitly specify any ring atom names (CENT/X is also available, of course).
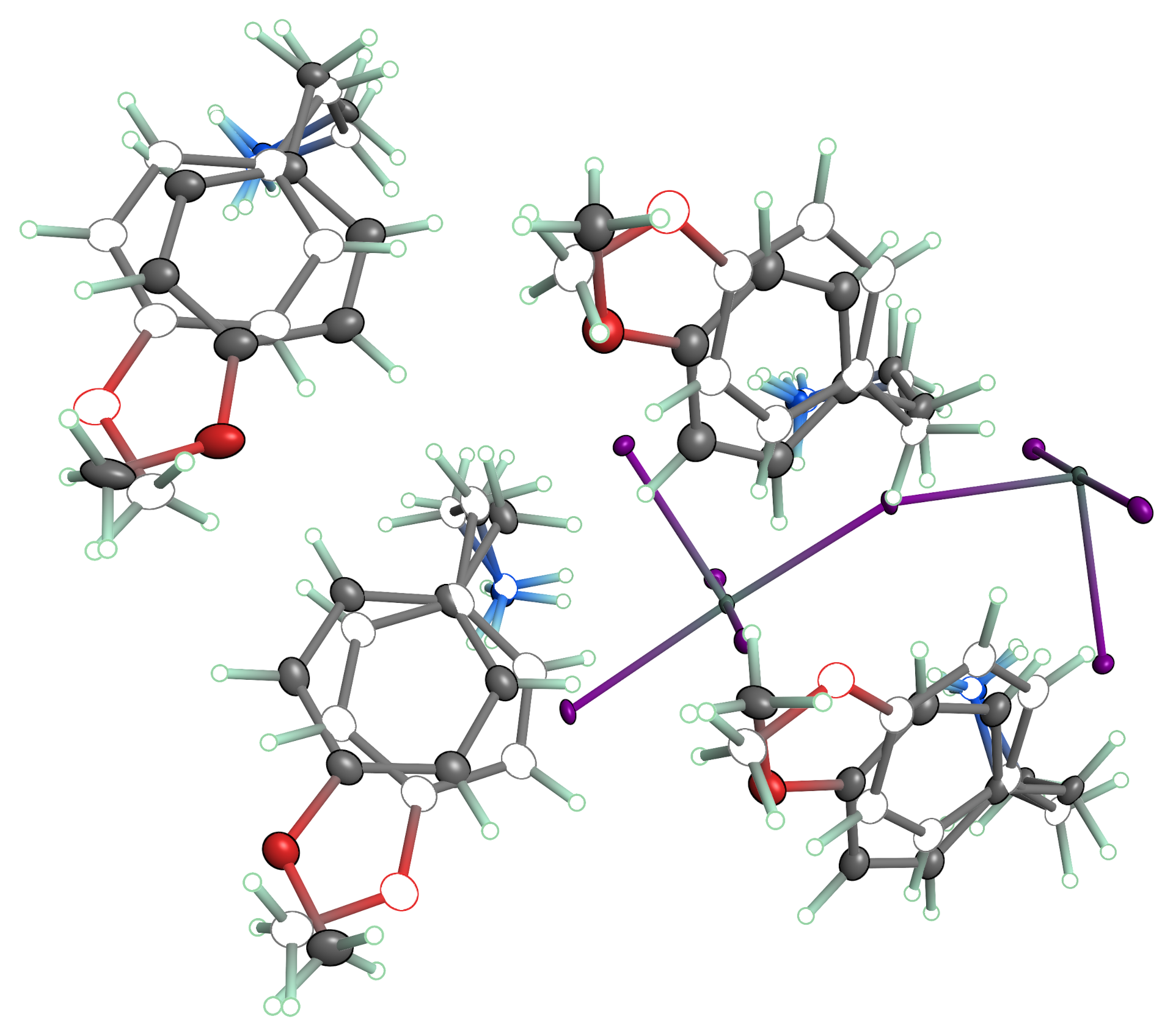
Disorder visualization in PrimeXP is flexible and can be shown in a number of ways. One such view is given above, in which major occupancy groups are drawn with 3D shading and minor disorder moieties are shown as boundary ellipses. Note also the colour blending in the bonds. A click button cycles through split-coloured bonds, blended bonds, and grey/black (depending on the scene).
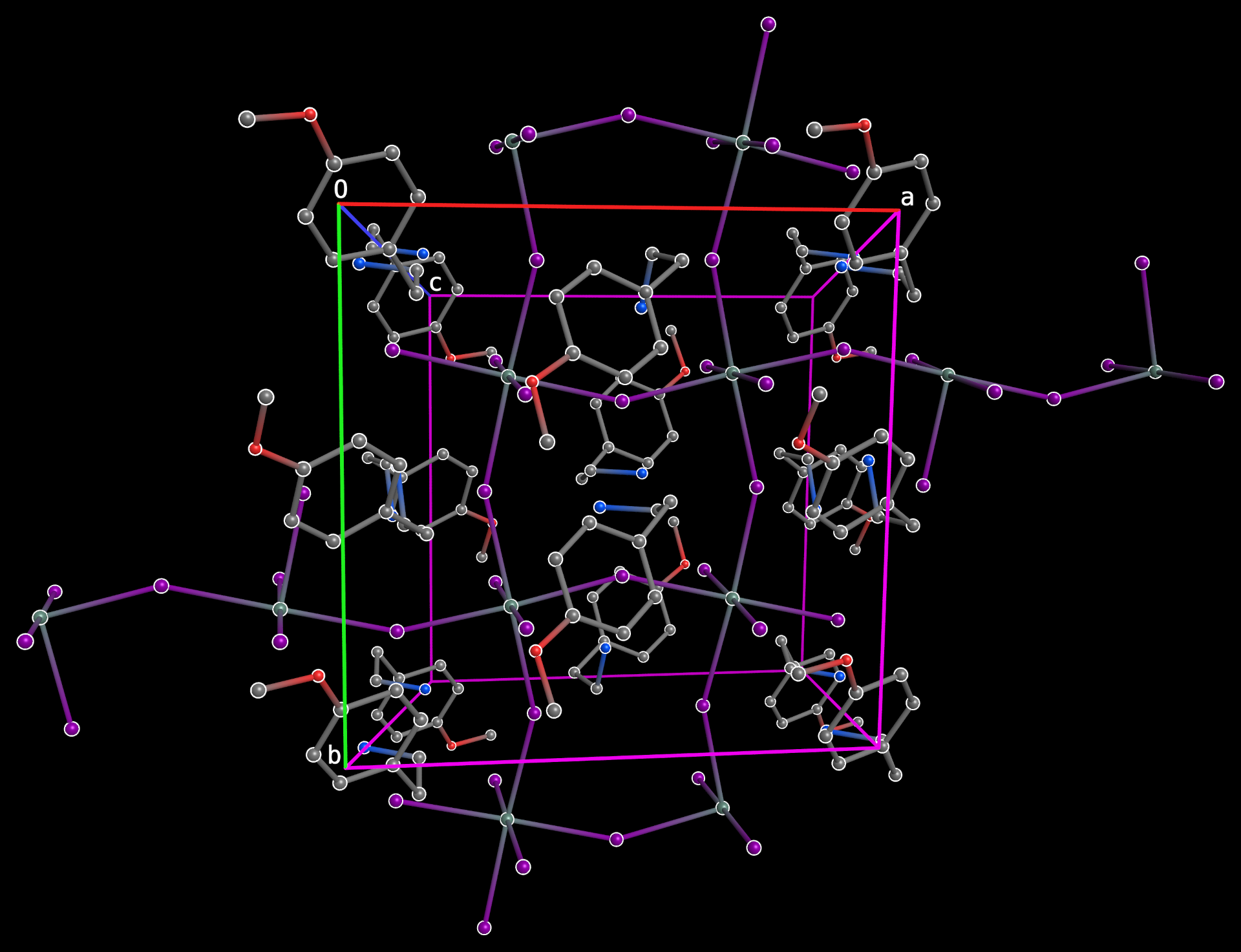
There are also many ways to construct packing plots in PrimeXP. The above shows a perspective view down the c-axis of a di-periodic perovskite-like structure (same structure as the disorder plot (vide supra ), but with minor disorder removed). Orthogonal projections are also available at the click of an on-screen button.
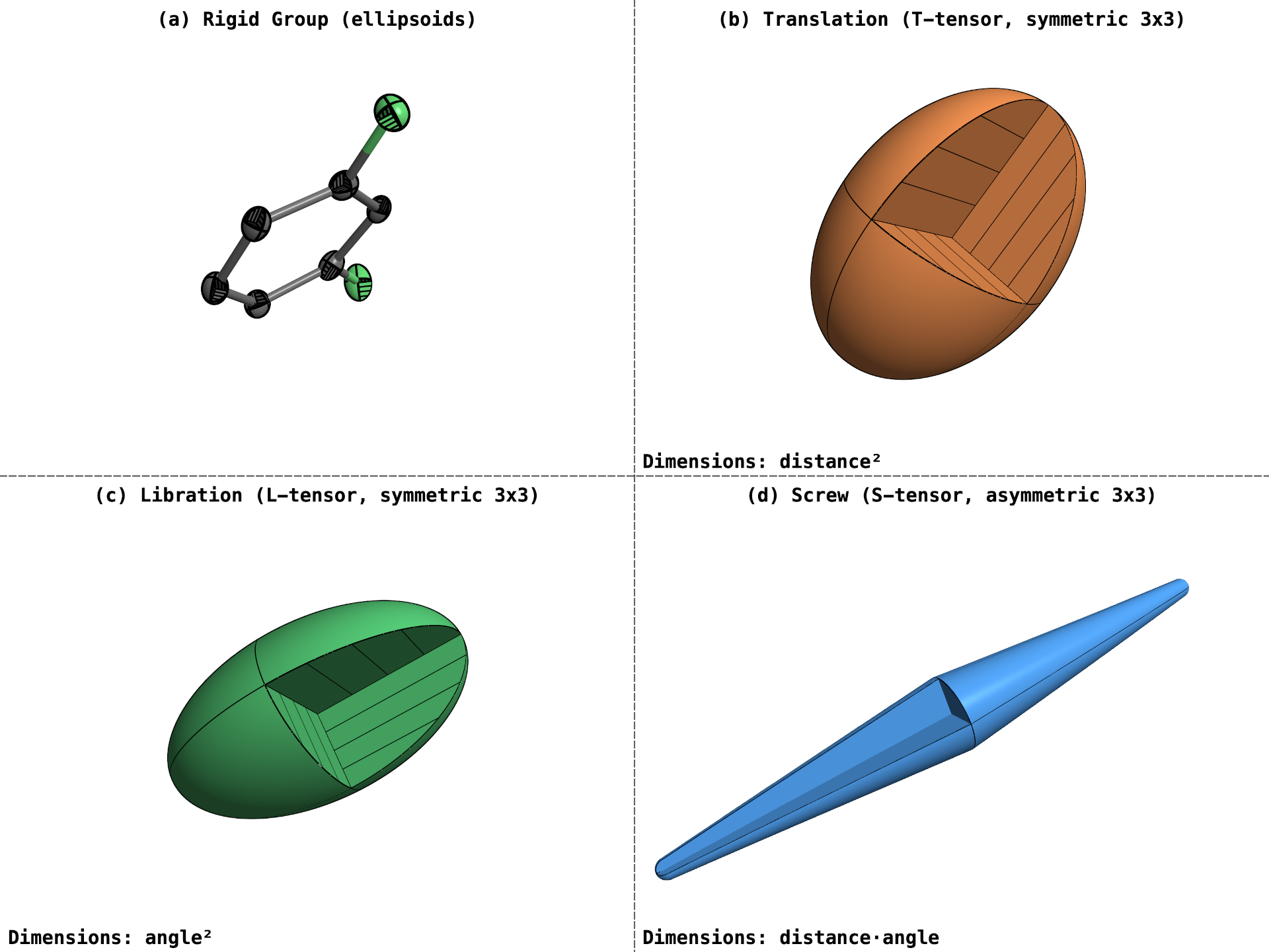
Real-time translation-libration-screw (TLS) visualisation. The translation (T) and libration (rotary oscillation, L) tensors can be represented by 3x3 symmetric positive-definite matrices, and so may be visualized as ellipsoids, similar to atomic displacement ellipsoids. The screw (S) tensor on the other hand, represents the coupling of translation and rotation. This requires an asymmetric 3x3 matrix, which can't be drawn as an ellipsoid. In PrimeXP, the S-tensor is visualized as an elongated round-ended bicone. The long axis of the bicone corresponds to the direction of screw propagation, and is wider in the middle where atomic movement is greatest.
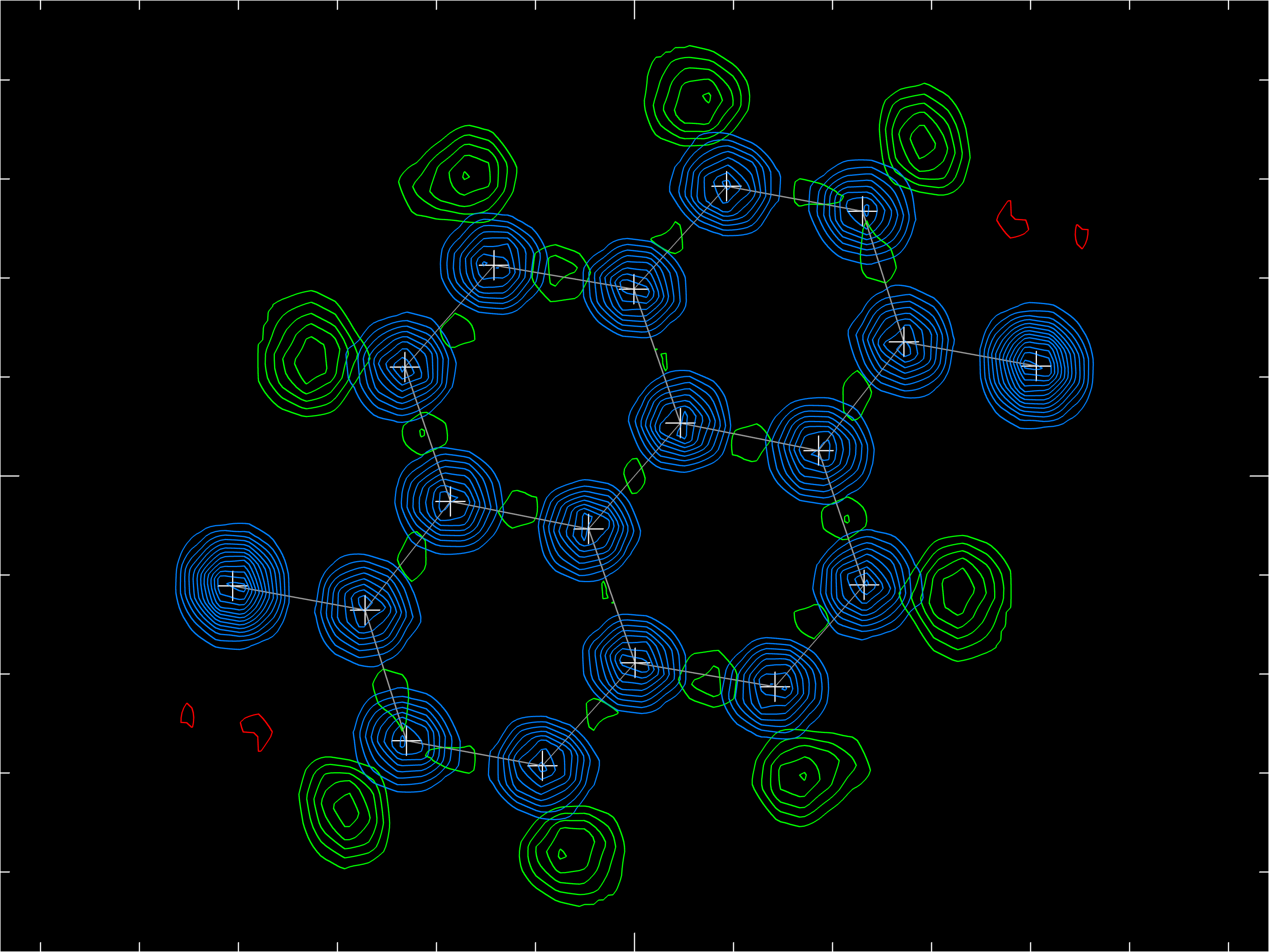
Old-style 2D electron-density contour plots similar those made by the EDEN command inXP are possible if a *.fcf file in 'LIST 6' format is available to PrimeXP 's version of EDEN. In the above, the blue contours represent experimental electron density for the carbon and fluorine atoms. The model did not have hydrogen atoms included, so they show up as prominent positive contours in the difference map (green) at the expected positions. The small red contours indicate where the difference map happened to be negative. This view may be zoomed in and out by mouse action.
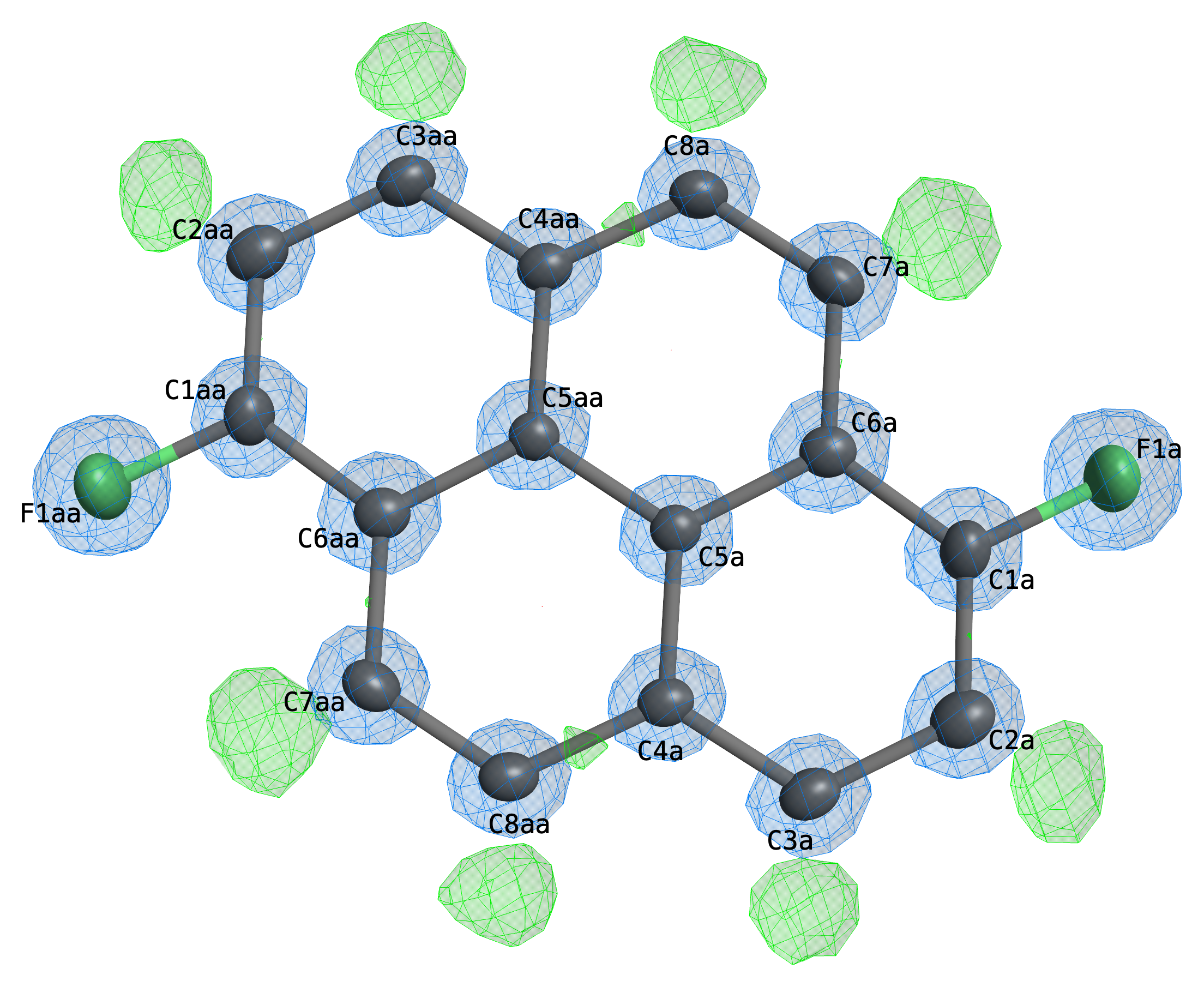
Probably more useful nowadays than 2D electron-density contours are 3D representations. Above is such a plot of the same molecule, again showing observed density in blue and difference density in green. In 3D, the electron density may be zoomed in/out and rotated in real time. The EDEN command also creates a conditional four-state click button on the graphical display that cycle through different visualisation styles (cage contours, diffuse clouds, clouds and cages superimposed (as above), and off).
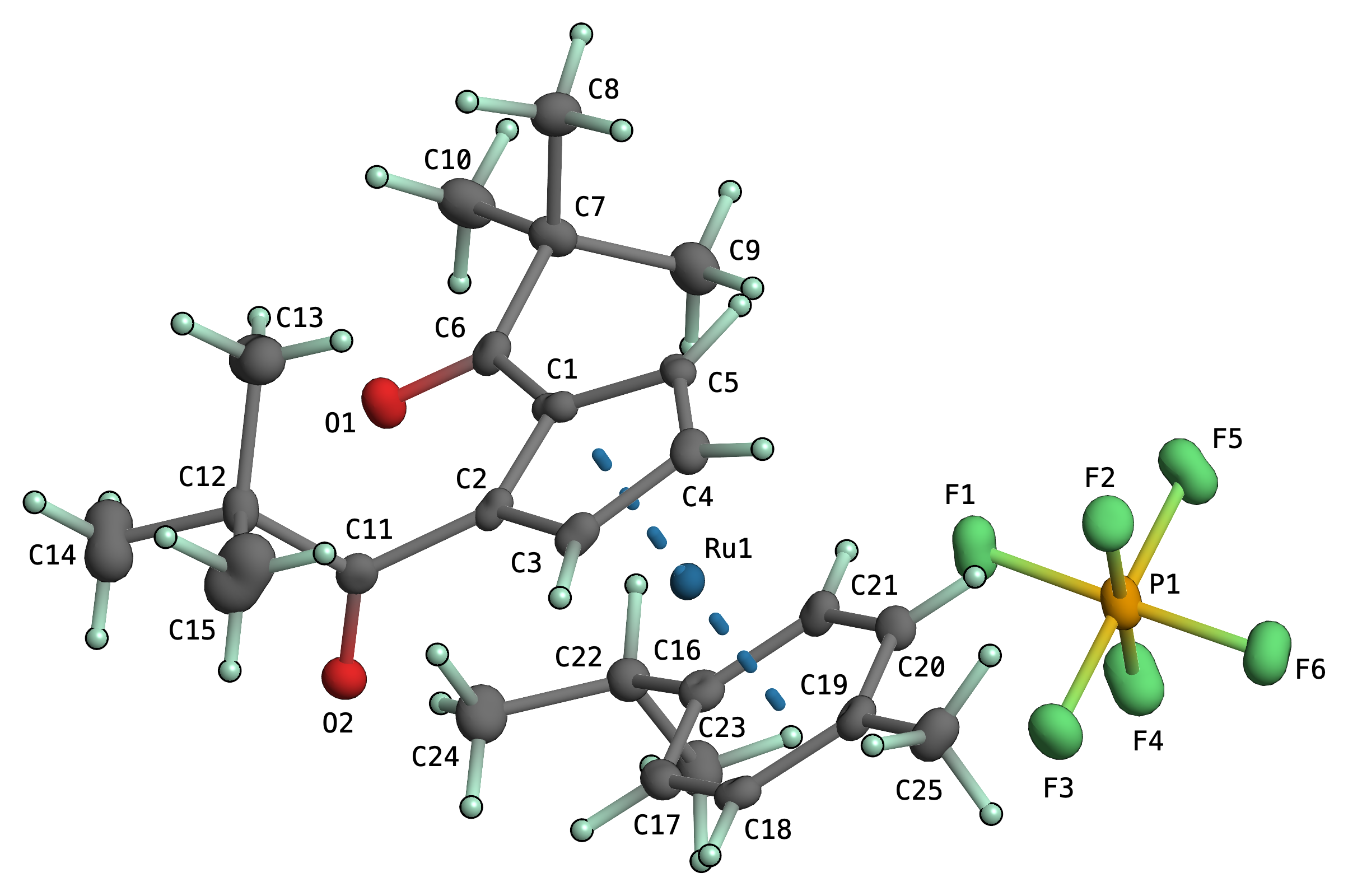
Anisotropic displacement parameter plots in the style of Bürgi's PEANUT program are now possible in primeXP. Both RMSD (root-mean-square-displacement) and MSD (mean-square-displacement), as either mesh-contours, 3D solid rendered (as above), or both (mesh superimposed on 3D solid) are possible. These are fully 3D interactive and can be freely mixed with other atom visualization styles, globally, by group selection, or on a per-atom basis. Difference 'peanuts' might follow.
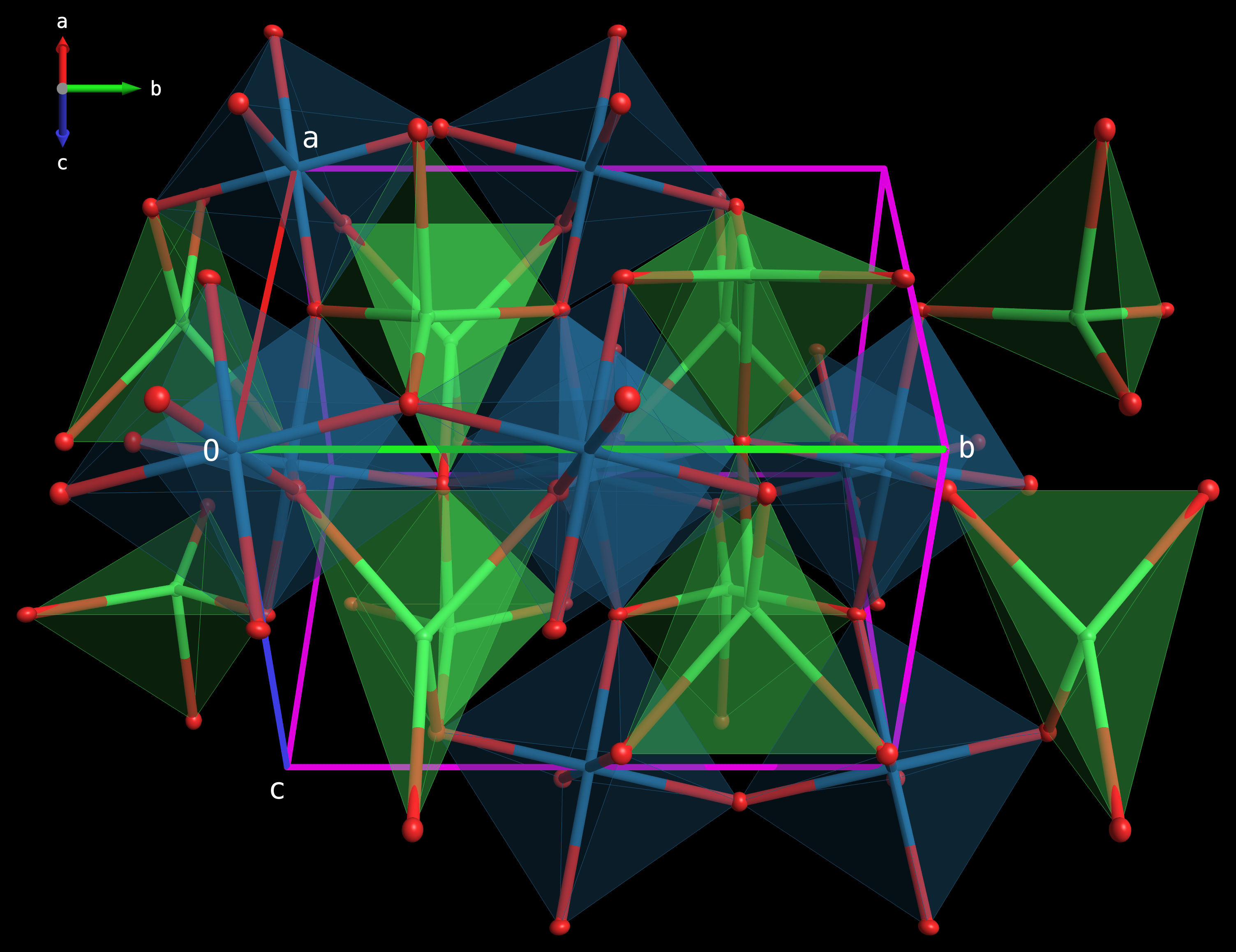
Graphical representations of coordination geometries by polyhedral convex hulls around selected central atoms are now implemented. The program identifies bonded ligands, calculates the 3D faces connecting them, and reports the coordination number (CN), number of faces, and calculated volume for each generated polyhedron. The resulting polyhedra are rendered with semi-transparent faces and wireframe edges along with the atoms and bonds present in the structure. As with all other 3D view modes, these are fully rotatable and zoomable via mouse action.
Availability and Implementation
The modular architecture of PrimeXP separates graphical routines, command parsing, and crystallographic symmetry and mathematics. It was developed on macOS, and runs with a few minor modifications to some screen-resolution settings on Linux (thanks to Paul Boyle). It should also run on Windows, but that has not yet been tried because I don't use Windows unless I have to, which in retirement, I don't. Although PrimeXP will probably always be a work in progress, and new features are still being added, the program is close to being ready for general release. Once a few remaining known bugs etc. have been fixed, I'll make it freely available.